1. Introduction¶
[ ]:
# Install CuPy and import as np !
!curl https://colab.chainer.org/install | sh -
import cupy as np
import numpy
Reading package lists... Done
Building dependency tree
Reading state information... Done
libcusparse8.0 is already the newest version (8.0.61-1).
libnvrtc8.0 is already the newest version (8.0.61-1).
libnvtoolsext1 is already the newest version (8.0.61-1).
0 upgraded, 0 newly installed, 0 to remove and 0 not upgraded.
Requirement already satisfied: cupy-cuda80==4.0.0b3 from https://github.com/kmaehashi/chainer-colab/releases/download/2018-02-06/cupy_cuda80-4.0.0b3-cp36-cp36m-linux_x86_64.whl in /usr/local/lib/python3.6/dist-packages
Requirement already satisfied: fastrlock>=0.3 in /usr/local/lib/python3.6/dist-packages (from cupy-cuda80==4.0.0b3)
Requirement already satisfied: six>=1.9.0 in /usr/local/lib/python3.6/dist-packages (from cupy-cuda80==4.0.0b3)
Requirement already satisfied: numpy>=1.9.0 in /usr/local/lib/python3.6/dist-packages (from cupy-cuda80==4.0.0b3)
[ ]:
class Regressor(object):
"""
Base class for regressors
"""
def fit(self, X, t, **kwargs):
"""
estimates parameters given training dataset
Parameters
----------
X : (sample_size, n_features) np.ndarray
training data input
t : (sample_size,) np.ndarray
training data target
"""
self._check_input(X)
self._check_target(t)
if hasattr(self, "_fit"):
self._fit(X, t, **kwargs)
else:
raise NotImplementedError
def predict(self, X, **kwargs):
"""
predict outputs of the model
Parameters
----------
X : (sample_size, n_features) np.ndarray
samples to predict their output
Returns
-------
y : (sample_size,) np.ndarray
prediction of each sample
"""
self._check_input(X)
if hasattr(self, "_predict"):
return self._predict(X, **kwargs)
else:
raise NotImplementedError
def _check_input(self, X):
if not isinstance(X, np.ndarray):
raise ValueError("X(input) is not np.ndarray")
if X.ndim != 2:
raise ValueError("X(input) is not two dimensional array")
if hasattr(self, "n_features") and self.n_features != np.size(X, 1):
raise ValueError(
"mismatch in dimension 1 of X(input) "
"(size {} is different from {})"
.format(np.size(X, 1), self.n_features)
)
def _check_target(self, t):
if not isinstance(t, np.ndarray):
raise ValueError("t(target) must be np.ndarray")
if t.ndim != 1:
raise ValueError("t(target) must be one dimenional array")
[ ]:
class LinearRegressor(Regressor):
"""
Linear regression model
y = X @ w
t ~ N(t|X @ w, var)
"""
def _fit(self, X, t):
self.w = np.linalg.pinv(X) @ t
self.var = np.mean(np.square(X @ self.w - t))
def _predict(self, X, return_std=False):
y = X @ self.w
if return_std:
y_std = np.sqrt(self.var) + np.zeros_like(y)
return y, y_std
return y
[ ]:
import matplotlib.pyplot as plt
%matplotlib inline
np.random.seed(1234)
1.1. Example: Polynomial Curve Fitting¶
[ ]:
def create_toy_data(func, sample_size, std):
x = np.linspace(0, 1, sample_size)
t = func(x) + np.random.normal(scale=std, size=x.shape)
return x, t
def func(x):
return np.sin(2 * 3.14 * x) # 3.14 should be np.pi
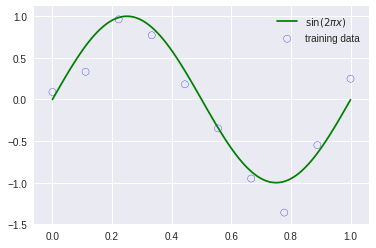
x_train, y_train = create_toy_data(func, 10, 0.25)
x_test = np.linspace(0, 1, 100)
y_test = func(x_test)
plt.scatter(x_train.get(), y_train.get(), facecolor="none", edgecolor="b", s=50, label="training data")
plt.plot(x_test.get(), y_test.get(), c="g", label="$\sin(2\pi x)$")
plt.legend()
plt.show()
[ ]:
import itertools
import functools
class PolynomialFeatures(object):
"""
polynomial features
transforms input array with polynomial features
Example
=======
x =
[[a, b],
[c, d]]
y = PolynomialFeatures(degree=2).transform(x)
y =
[[1, a, b, a^2, a * b, b^2],
[1, c, d, c^2, c * d, d^2]]
"""
def __init__(self, degree=2):
"""
construct polynomial features
Parameters
----------
degree : int
degree of polynomial
"""
assert isinstance(degree, int)
self.degree = degree
def transform(self, x):
"""
transforms input array with polynomial features
Parameters
----------
x : (sample_size, n) ndarray
input array
Returns
-------
output : (sample_size, 1 + nC1 + ... + nCd) ndarray
polynomial features
"""
if x.ndim == 1:
x = x[:, None]
x_t = x.transpose().get() # https://github.com/cupy/cupy/issues/1084
features = [numpy.ones(len(x))] # https://github.com/cupy/cupy/issues/1084
for degree in range(1, self.degree + 1):
for items in itertools.combinations_with_replacement(x_t, degree):
features.append(functools.reduce(lambda x, y: x * y, items))
features = numpy.array(features) # https://github.com/cupy/cupy/issues/1084
return np.asarray(features).transpose()
[ ]:
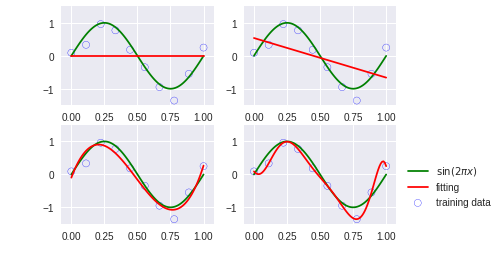
for i, degree in enumerate([0, 1, 3, 9]):
plt.subplot(2, 2, i + 1)
feature = PolynomialFeatures(degree)
X_train = feature.transform(x_train)
X_test = feature.transform(x_test)
model = LinearRegressor()
model.fit(X_train, y_train)
y = model.predict(X_test)
plt.scatter(x_train.get(), y_train.get(), facecolor="none", edgecolor="b", s=50, label="training data")
plt.plot(x_test.get(), y_test.get(), c="g", label="$\sin(2\pi x)$")
plt.plot(x_test.get(), y.get(), c="r", label="fitting")
plt.ylim(-1.5, 1.5)
plt.annotate("M={}".format(degree), xy=(-0.15, 1))
plt.legend(bbox_to_anchor=(1.05, 0.64), loc=2, borderaxespad=0.)
plt.show()
[ ]:
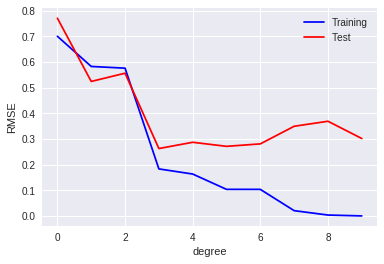
def rmse(a, b):
return np.sqrt(np.mean(np.square(a - b)))
training_errors = []
test_errors = []
for i in range(10):
feature = PolynomialFeatures(i)
X_train = feature.transform(x_train)
X_test = feature.transform(x_test)
model = LinearRegressor()
model.fit(X_train, y_train)
y = model.predict(X_test)
training_errors.append(rmse(model.predict(X_train), y_train))
test_errors.append(rmse(model.predict(X_test), y_test + np.random.normal(scale=0.25, size=len(y_test))))
plt.plot(training_errors, 'o-', mfc="none", mec="b", ms=10, c="b", label="Training")
plt.plot(test_errors, 'o-', mfc="none", mec="r", ms=10, c="r", label="Test")
plt.legend()
plt.xlabel("degree")
plt.ylabel("RMSE")
plt.show()
Regularization¶
[ ]:
class RidgeRegressor(Regressor):
"""
Ridge regression model
w* = argmin |t - X @ w| + a * |w|_2^2
"""
def __init__(self, alpha=1.):
self.alpha = alpha
def _fit(self, X, t):
eye = np.eye(np.size(X, 1))
self.w = np.linalg.solve(self.alpha * eye + X.T @ X, X.T @ t)
def _predict(self, X):
y = X @ self.w
return y
[ ]:
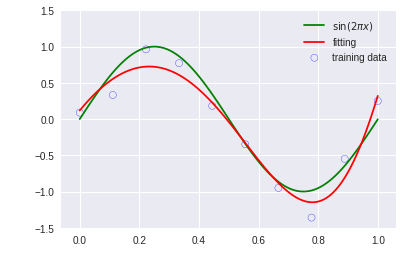
feature = PolynomialFeatures(9)
X_train = feature.transform(x_train)
X_test = feature.transform(x_test)
model = RidgeRegressor(alpha=1e-3)
model.fit(X_train, y_train)
y = model.predict(X_test)
y = model.predict(X_test)
plt.scatter(x_train.get(), y_train.get(), facecolor="none", edgecolor="b", s=50, label="training data")
plt.plot(x_test.get(), y_test.get(), c="g", label="$\sin(2\pi x)$")
plt.plot(x_test.get(), y.get(), c="r", label="fitting")
plt.ylim(-1.5, 1.5)
plt.legend()
plt.annotate("M=9", xy=(-0.15, 1))
plt.show()
1.2.6 Bayesian curve fitting¶
[ ]:
class BayesianRegressor(Regressor):
"""
Bayesian regression model
w ~ N(w|0, alpha^(-1)I)
y = X @ w
t ~ N(t|X @ w, beta^(-1))
"""
def __init__(self, alpha=1., beta=1.):
self.alpha = alpha
self.beta = beta
self.w_mean = None
self.w_precision = None
def _fit(self, X, t):
if self.w_mean is not None:
mean_prev = self.w_mean
else:
mean_prev = np.zeros(np.size(X, 1))
if self.w_precision is not None:
precision_prev = self.w_precision
else:
precision_prev = self.alpha * np.eye(np.size(X, 1))
w_precision = precision_prev + self.beta * X.T @ X
w_mean = np.linalg.solve(
w_precision,
precision_prev @ mean_prev + self.beta * X.T @ t
)
self.w_mean = w_mean
self.w_precision = w_precision
self.w_cov = np.linalg.inv(self.w_precision)
def _predict(self, X, return_std=False, sample_size=None):
if isinstance(sample_size, int):
w_sample = np.random.multivariate_normal(
self.w_mean, self.w_cov, size=sample_size
)
y = X @ w_sample.T
return y
y = X @ self.w_mean
if return_std:
y_var = 1 / self.beta + np.sum(X @ self.w_cov * X, axis=1)
y_std = np.sqrt(y_var)
return y, y_std
return y
[ ]:
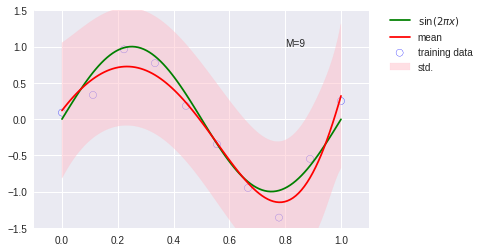
model = BayesianRegressor(alpha=2e-3, beta=2)
model.fit(X_train, y_train)
y, y_err = model.predict(X_test, return_std=True)
plt.scatter(x_train.get(), y_train.get(), facecolor="none", edgecolor="b", s=50, label="training data")
plt.plot(x_test.get(), y_test.get(), c="g", label="$\sin(2\pi x)$")
plt.plot(x_test.get(), y.get(), c="r", label="mean")
plt.fill_between(x_test.get(), (y - y_err).get(), (y + y_err).get(), color="pink", label="std.", alpha=0.5)
plt.xlim(-0.1, 1.1)
plt.ylim(-1.5, 1.5)
plt.annotate("M=9", xy=(0.8, 1))
plt.legend(bbox_to_anchor=(1.05, 1.), loc=2, borderaxespad=0.)
plt.show()